Overview
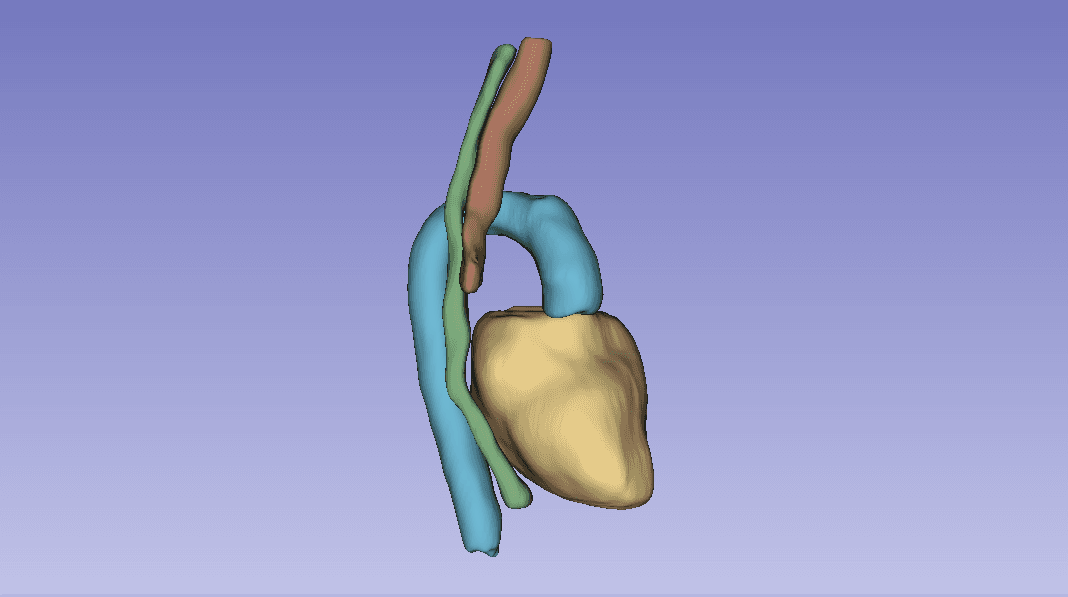
Group project for AI for Medical Imaging (UvA, part of the VU M.Sc. AI program) on the SegTHOR challenge: segmenting four thoracic organs (heart, esophagus, trachea, aorta) from 3D CT volumes. We compared a 2D baseline (ENet), a custom 2D architecture (CustomNet, U-Net based), and 3D Full-Resolution nnU-Net.
A real data problem first
Before any modelling, the heart annotations were misaligned for several patients. Patient 27 had two ground truth files in the dataset, one correct and one misaligned. We tried to fix the misalignment with Elastix (rigid + affine transformations), but no parameter combination got the heart to register cleanly.
The fix that worked was a centroid-then-rotation approach: subtract the misaligned heart's centroid from the correct centroid to get a translation vector, then sweep rotation angles in the transverse plane and pick the one that maximised IoU. The optimal correction was a −26.6° rotation around the superior-inferior axis. We then applied the same transformation across the dataset.
This took longer than it should have and is the project's biggest concrete lesson: spend time on the data before architecture experiments.
Architectures and training
- ENet (2D baseline). Lightweight encoder-decoder with bottleneck residual blocks. Slice-wise prediction.
- CustomNet (2D). U-Net derivative; encoder filters grow 64 → 512 with DoubleConv blocks and ReLU. AdamW (lr 1e-3, β₁ 0.9, β₂ 0.999, weight decay 1e-4), 100 epochs.
- nnU-Net (3D Full Resolution). Self-configuring framework that picks its own preprocessing, architecture, and training settings for the dataset.
Loss: combined Dice + cross-entropy with 0.7 / 0.3 weighting to handle the heart-vs-esophagus class imbalance, plus class-specific weights inside the Dice term (esophagus weighted 0.3, background 0.1, others 0.2).
Metrics: Dice coefficient, Hausdorff distance, ASSD (average symmetric surface distance). Hausdorff alone is outlier-prone, so we used both Hausdorff and ASSD together.
Results
Comparisons use paired t-tests where the differences were normally distributed, and Mann-Whitney U where they weren't.
- CustomNet beat ENet on average Dice, t(78) = 2.10, p < 0.05. ASSD also improved significantly, U = 539.0, p < 0.05. Hausdorff trended down but the test was not significant (p = 0.08).
- nnU-Net beat both. vs ENet: Dice t(78) = 5.03, p < 0.01; recall t(78) = 5.68, p < 0.01. vs CustomNet: Dice t(78) = 3.42, p < 0.01; recall t(78) = 4.27, p < 0.01.
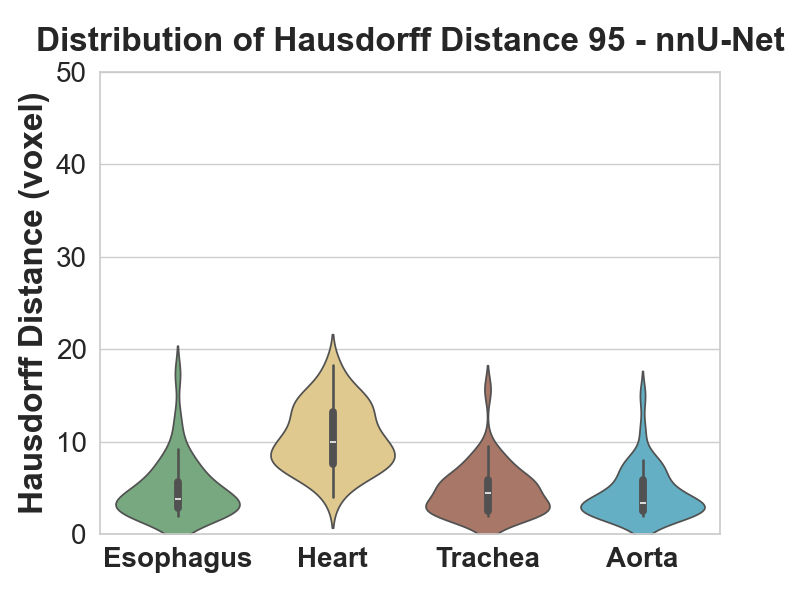
- The interesting nuance is the heart. Heart Hausdorff actually got worse for CustomNet despite a better Dice, because both CustomNet and nnU-Net over-segment the heart. Heart precision dropped from 0.92 (baseline) to 0.89 (CustomNet), and dropped further on nnU-Net. Recall went up; precision went down. Net Dice still improved but the failure mode is real.
- Where nnU-Net shines is the esophagus. The smallest, hardest organ. Dice distribution shifted up substantially, and both precision and recall improved, where the 2D models struggled most.
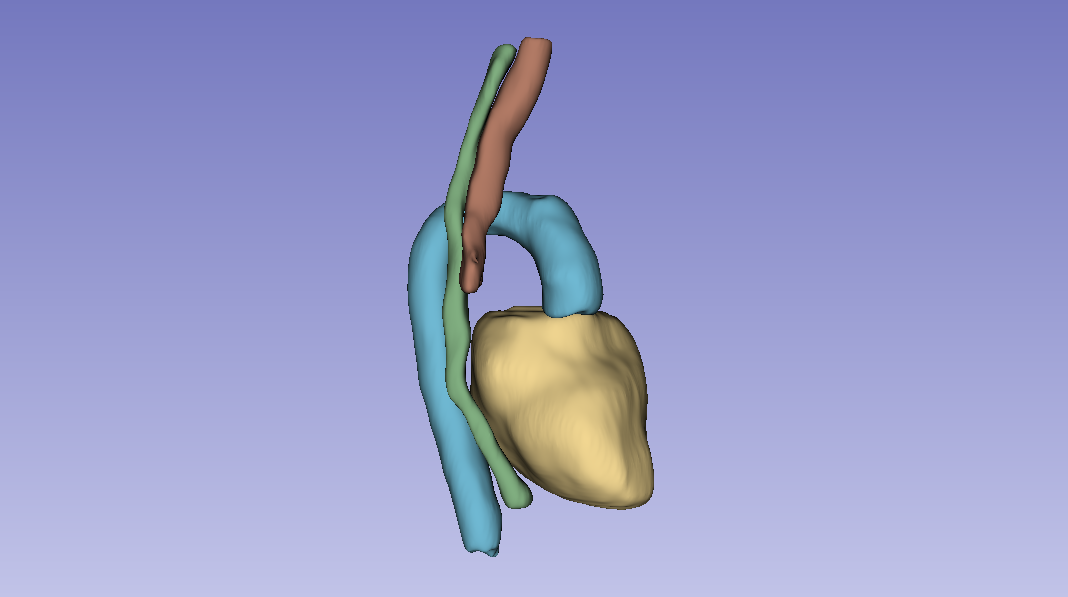
Takeaway
3D Full-Resolution nnU-Net is the clear winner for this task: significantly better Dice and recall, smoother predictions (post-processing's connected-component analysis cleans up the artefacts the 2D models leave). The cost is that it over-segments the heart, which would matter for clinical deployment and is the obvious next-step refinement.
Stack
Python, PyTorch, nnU-Net, ENet, Custom U-Net variant, Elastix (registration), Slicer, NIfTI, HPC (SLURM scheduler).